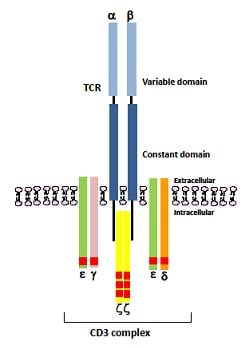
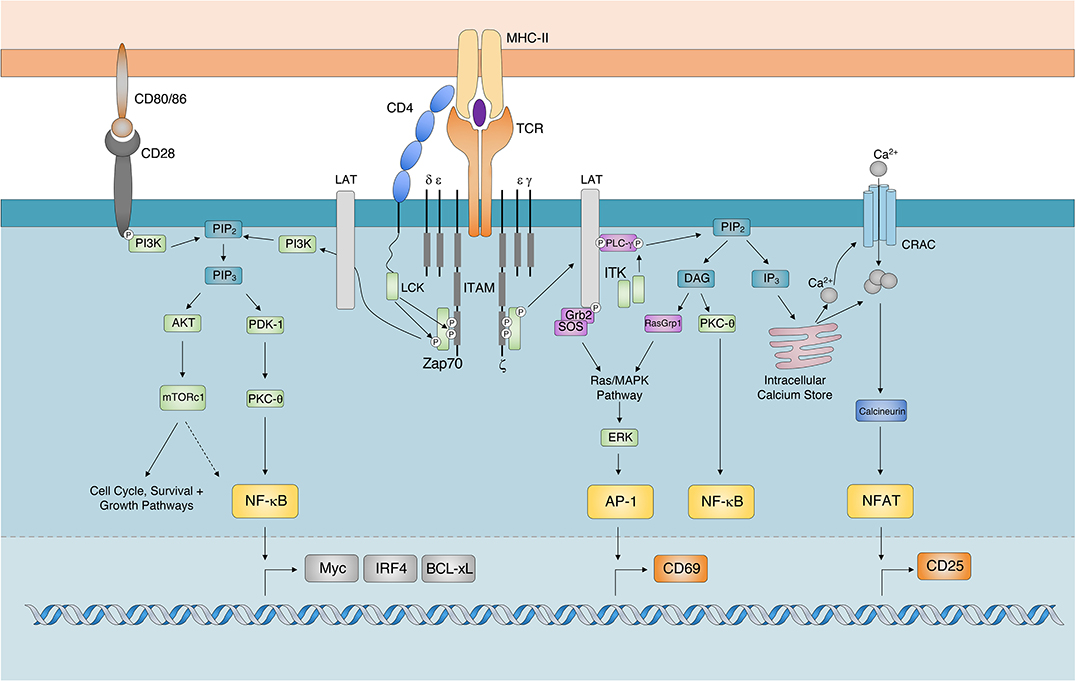
Isolation of TCR genes with tumor-killing activity from tumor-infiltrating and circulating lymphocytes in a tumor rejection cynomolgus macaque model: Molecular Therapy - Oncolytics
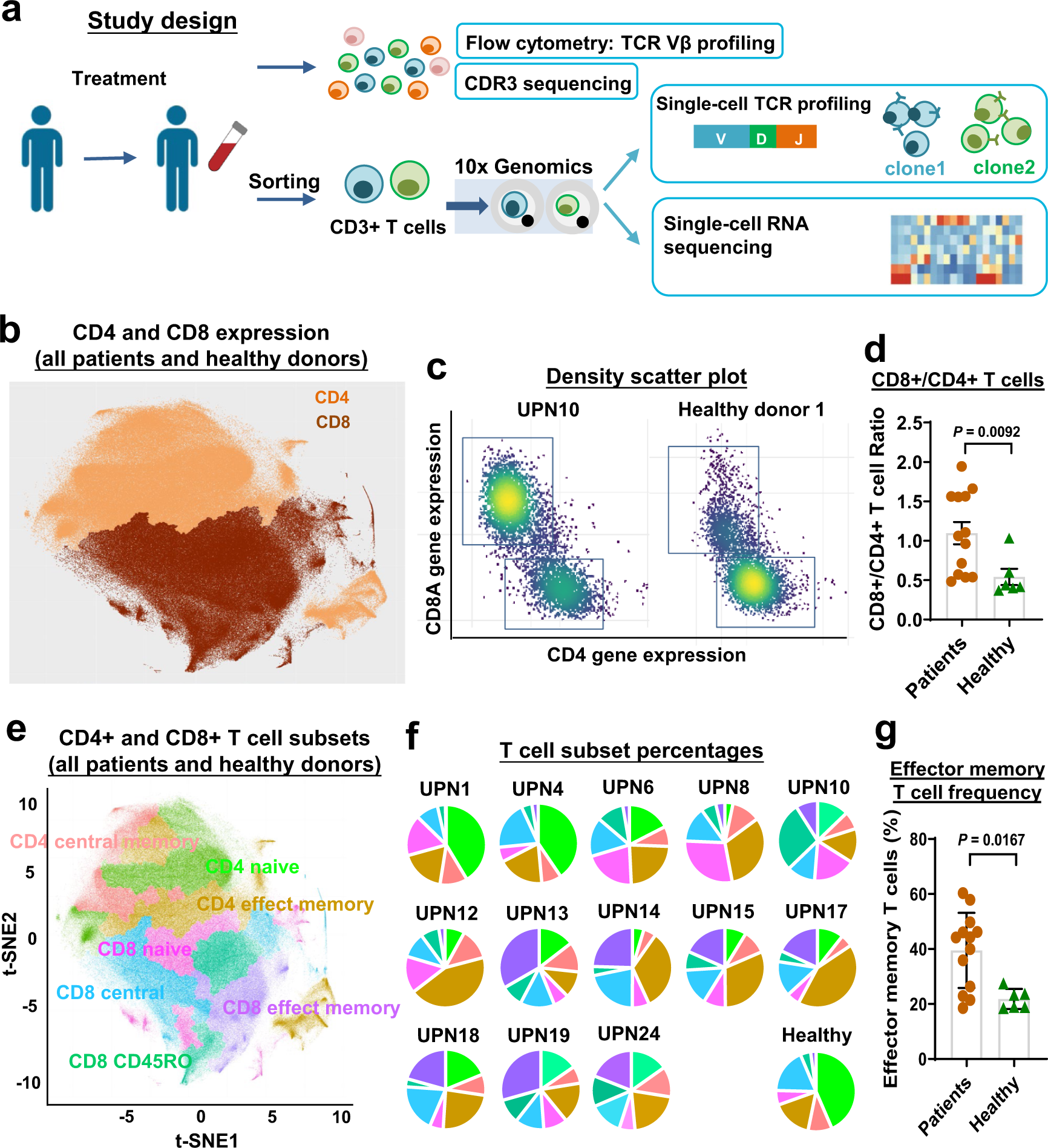
Single-cell RNA sequencing coupled to TCR profiling of large granular lymphocyte leukemia T cells | Nature Communications

Diversification of TCR Expression Level among TCR-iPSC Clones (A) A map... | Download Scientific Diagram

Autoreactive CD8 T cells in NOD mice exhibit phenotypic heterogeneity but restricted TCR gene usage | Life Science Alliance

Antigen and checkpoint receptor engagement recalibrates T cell receptor signal strength - ScienceDirect

A, FACS analysis of TCRb + CD44 and CD62L levels on gated CD4 or CD8 SP... | Download Scientific Diagram

Increased T cell receptor (TCR) signal strength induces expression of... | Download Scientific Diagram

High‐throughput single‐cell sequencing of paired TCRα and TCRβ genes for the direct expression‐cloning and functional analysis of murine T‐cell receptors - Ludwig - 2019 - European Journal of Immunology - Wiley Online

Ligand recognition by the γδ TCR and discrimination between homeostasis and stress conditions | Cellular & Molecular Immunology

Amazon.com : Teacher Created Resources Chalk Brights Liquid Chalk Markers (TCR20884) : Office Products
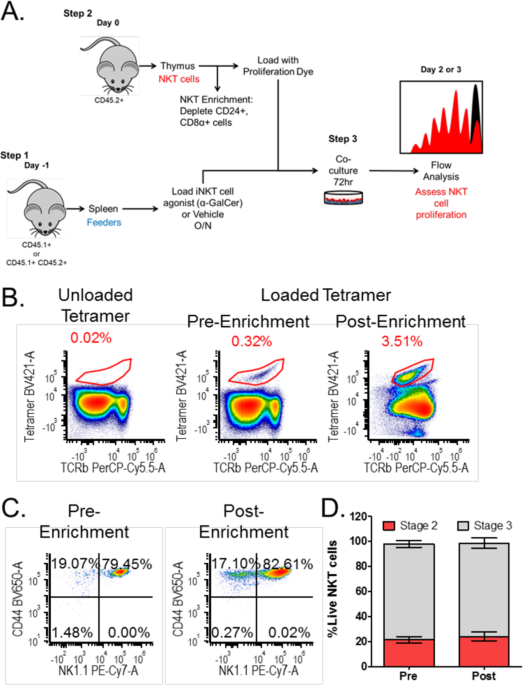
Thymic resident NKT cell subsets show differential requirements for CD28 co-stimulation during antigenic activation | Scientific Reports